diagnostics4u
LiverUSRecon: Automatic 3D Reconstruction and Volumetry of the Liver with a Few Partial Ultrasound Scans
MICCAI 2024 Paper (arXiv) Code
Authors
Kaushalya Sivayogaraj, Sahan T. Guruge, Udari Liyanage, Jeevani Udupihille, Saroj Jayasinghe, Gerard Fernando, Ranga Rodrigo, Rukshani Liyanaarachchi
📰 News
- Code is available at LiverUSRecon.
- To request access to the dataset, please contact Kaushalya Sivayogaraj.
- Pre-trained weights are available:
Abstract
3D reconstruction of the liver for volume measurement and 3D visual shape analysis using an accessible medical imaging modality like ultrasound (US) imaging is important. We present the first method capable of reconstructing the liver from a few partial ultrasound scans acquired at the midline, midclavicular line, and anterior-axillary line. To the best of our knowledge, this is the first automated deep learning method that calculates the liver volume from three incomplete 2D US scans. Further, we introduce a new US liver database with parallel, annotated CT scans comprising 134 scans. Our volumetry results are statistically closer to the ground-truth volumes obtained from CT scans than the volumes computed by radiologists using the Childs’ method.
Results
Ultrasound Segmentation and 3D Reconstruction
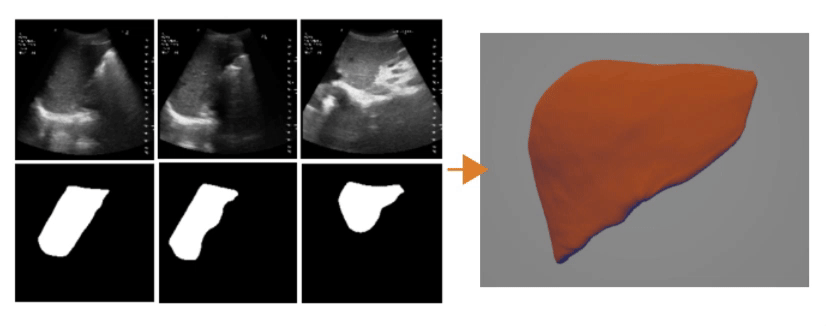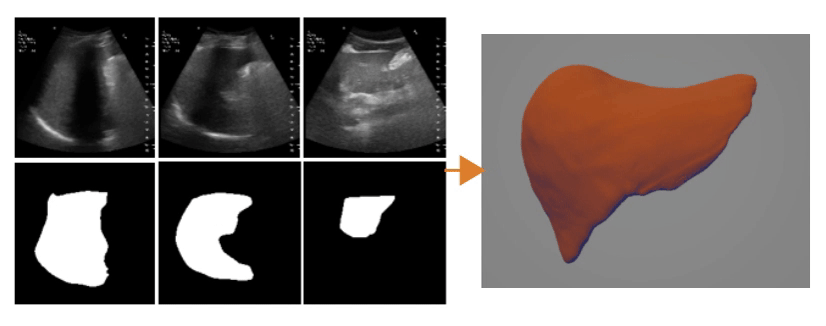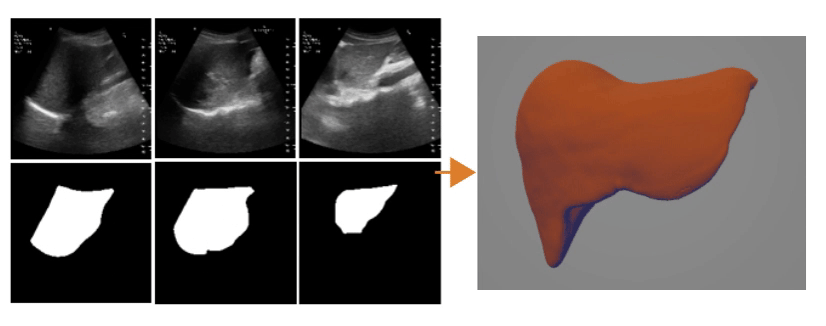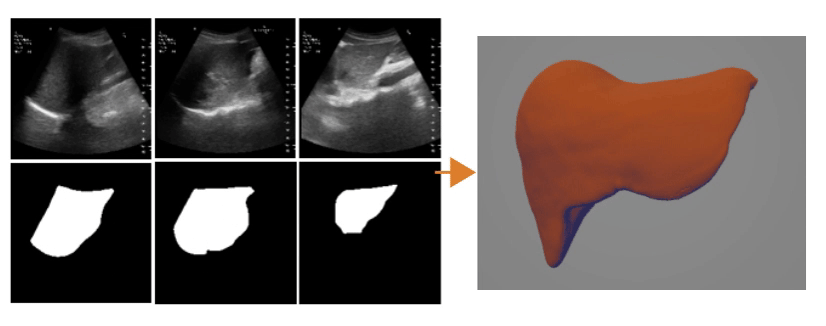
3D Reconstruction Overlap
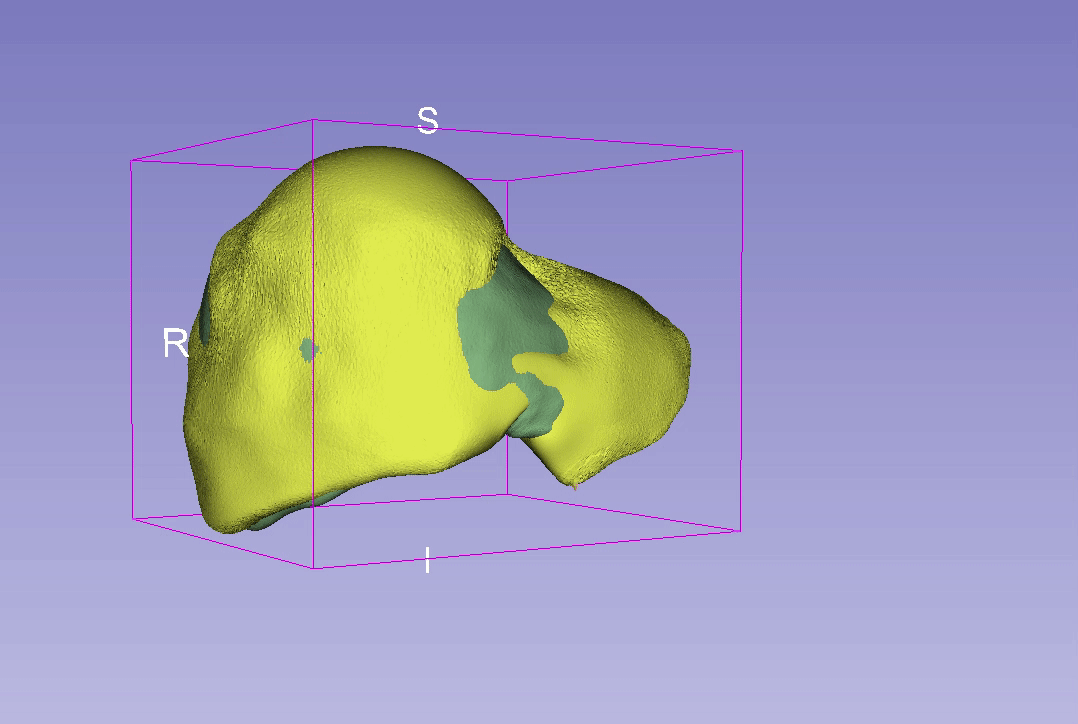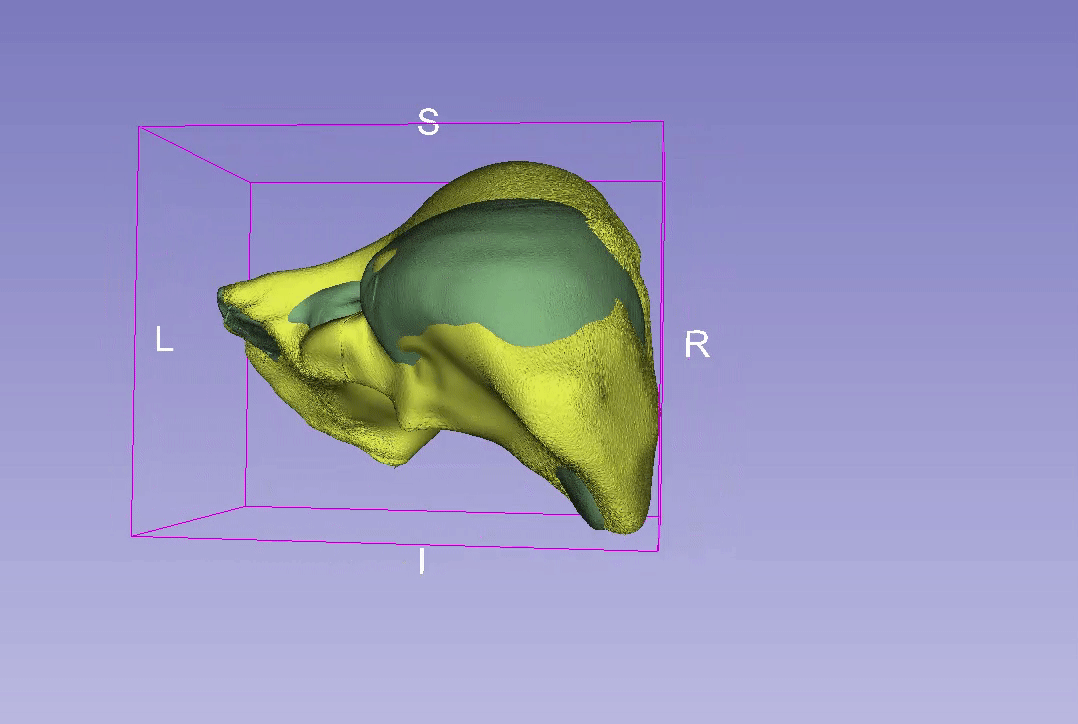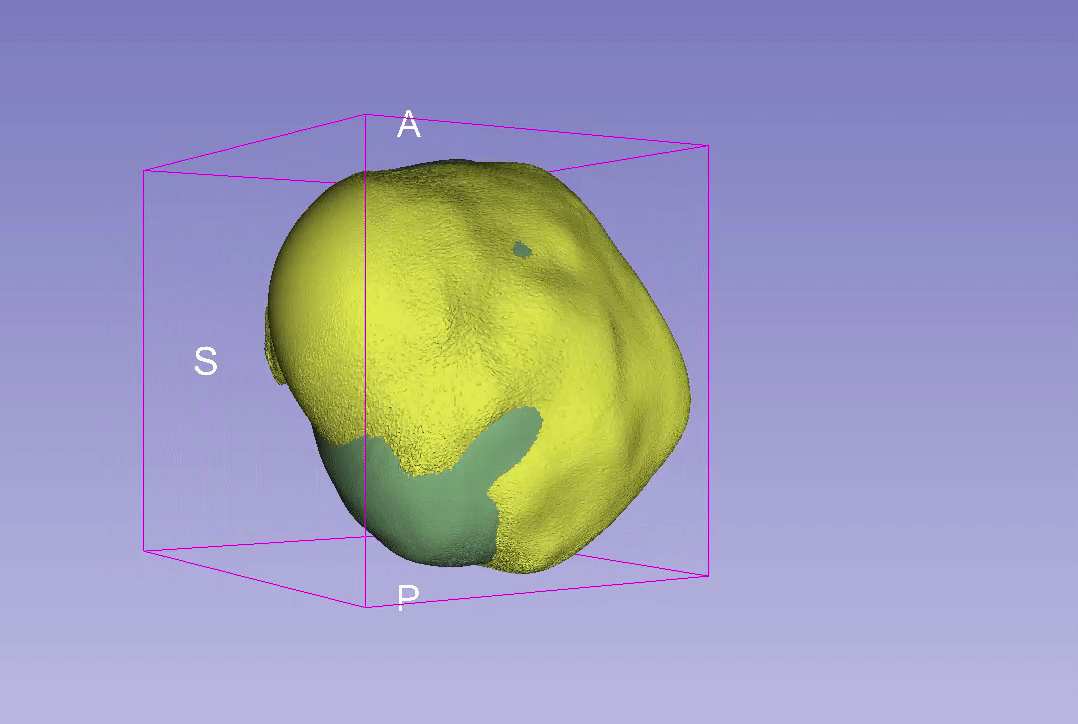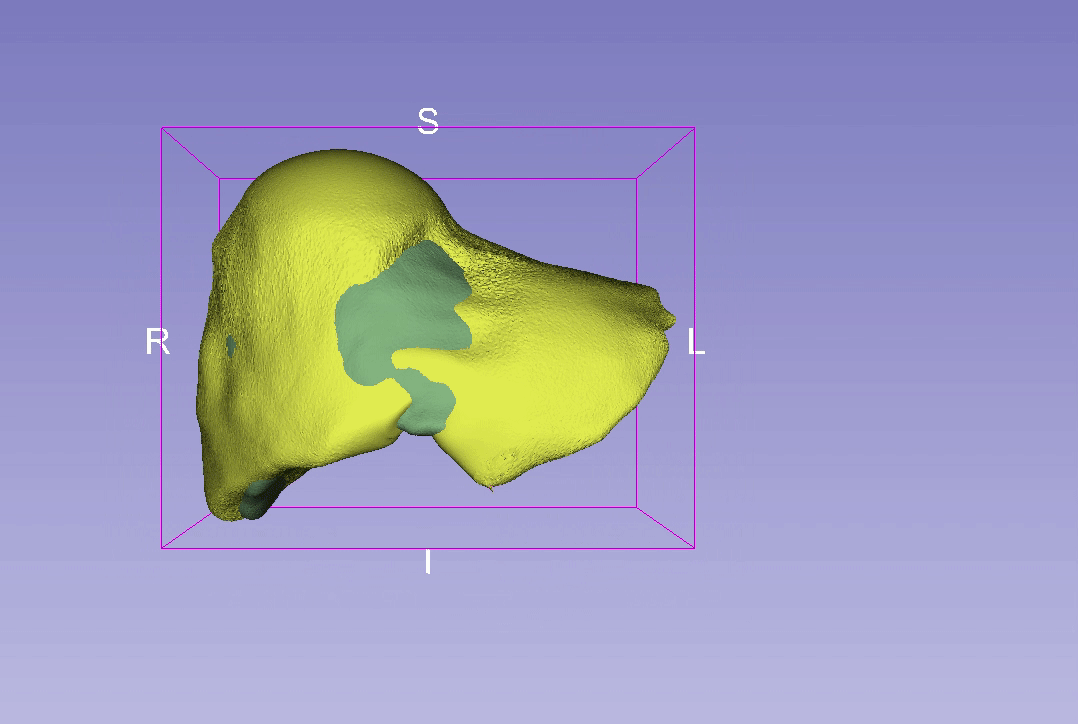
Point-to-Point Distance
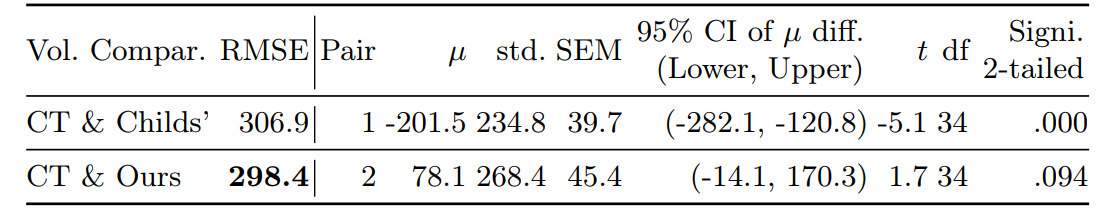
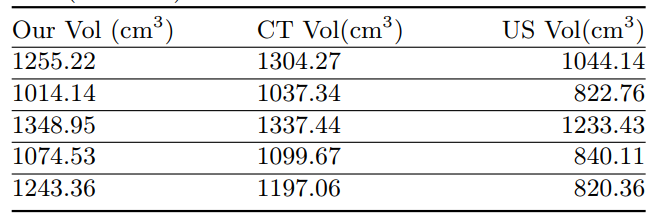
Statistical Analysis
Volume Comparison
Getting Started
1. Download Pre-trained ViT Model
- Download the R50-ViT-B_16 model from Google.
- Place the downloaded model in
./model/vit_checkpoint/imagenet21k/and rename it toR50-ViT-B_16.npz. - Download the pre-trained segmentation and reconstruction models:
- Place both models in a folder named
modelsunder the results directory.
2. Prepare the Dataset
- Contact Kaushalya Sivayogaraj to request access to the inference datasets.
3. Download Liver SSM Information
Download the following Statistical Shape Model (SSM) files and place them in ./SSM/:
| File | Link |
|---|---|
| Shape parameters | VT.txt |
| Mean shape | liver_aver.obj |
| PCA ratio | pca_ratio.txt |
| Normalization info | nor_list.txt |
4. Environment Setup
Create a Python 3.7 environment and install the required dependencies:
pip install -r requirements.txt
5. Inference
Run the inference script on the downloaded dataset:
CUDA_VISIBLE_DEVICES=0 python inference_liverusrecon.py \
--inference {dataset path} \
--save {results path} \
--ssm_info {ssm_info path}
Licenses
Code
Copyright © 2024 Zone24x7, Inc.
Code is licensed under the GNU Affero General Public License v3.0. You should have received a copy of the GNU Affero General Public License along with this code. If not, see https://www.gnu.org/licenses/.
ML Weights
Copyright © Zone24x7, Inc.
ML Weights are licensed under the Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. You should have received a copy of the license along with this work. If not, see https://creativecommons.org/licenses/by-nc-nd/3.0/.
Patient Data
Copyright © Zone24x7, Inc.
Patient data is licensed under the Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. You should have received a copy of the license along with this work. If not, see https://creativecommons.org/licenses/by-nc-nd/3.0/.
Citation
If you find this work useful, please consider citing:
@InProceedings{Siv_LiverUSRecon_MICCAI2024,
author = {Sivayogaraj, Kaushalya and Guruge, Sahan I. T. and Liyanage, Udari A. and
Udupihille, Jeevani J. and Jayasinghe, Saroj and Fernando, Gerard M. X. and
Rodrigo, Ranga and Liyanaarachchi, Rukshani},
title = ,
booktitle = {Proceedings of Medical Image Computing and Computer Assisted Intervention},
year = {2024},
pages = {436--445}
}